dat <- gsheet2tbl("https://docs.google.com/spreadsheets/d/1vYXB1Ag-ouLgo9nLIelP1V0hz-ki0f7p-aOCAkmuxKI/edit#gid=1058316481")
theme_set(theme_minimal_grid())Meta-analysis in Plant Pathology
Bibliographic info
Total number of pubs
nrow(dat)[1] 85dat |>
tabyl(article_type) article_type n percent
Original Article 79 0.92941176
Review 6 0.07058824Pubs per year
dat |>
filter(pub_year > 2010) |>
nrow()[1] 73Pub type by year
p1 <- dat %>%
tabyl(pub_year, article_type) %>%
pivot_longer(names_to = "Type",
values_to = "n", 2:3) %>%
ggplot(aes(pub_year, n, fill = Type))+
geom_col()+
scale_x_continuous(breaks = c(1999, 2001, 2004, 2007, 2010, 2013, 2016,
2019, 2022))+
theme(legend.position = "bottom",
panel.grid.major=element_line(colour="grey94"))+
scale_fill_viridis_d(option = "C")+
scale_y_continuous(n.breaks = 10)+
labs( x = "Publication year", y = "Number of publications")
p1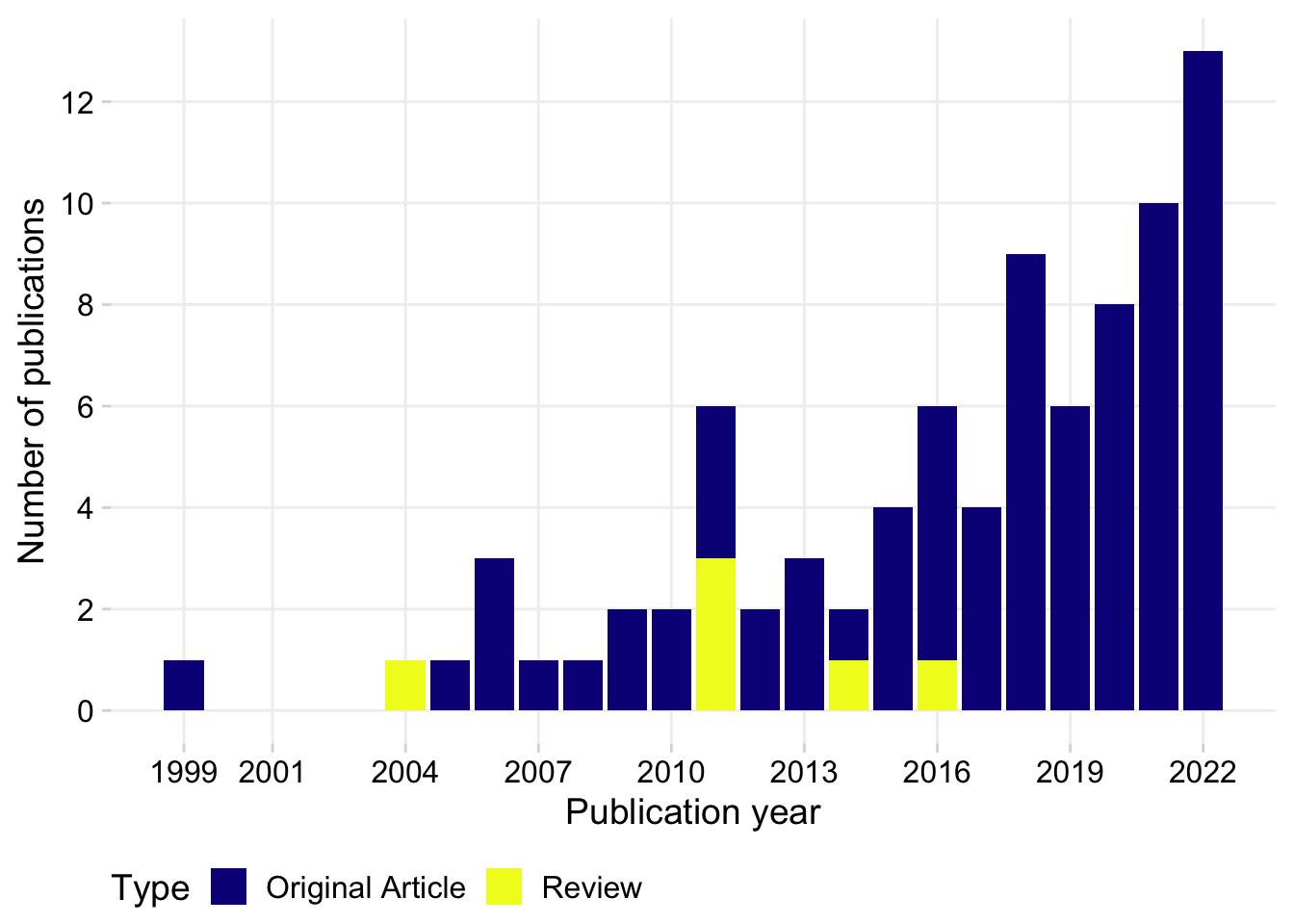
Journals
tab2 <- dat %>%
dplyr::select(journal) %>%
tabyl(journal) %>%
select(-percent) |>
arrange(-n)
tab2 journal n
Plant Disease 22
Phytopathology 21
Crop Protection 4
Plant Pathology 4
Biological Control 3
European Journal of Plant Pathology 3
Plant Health Progress 2
Summa Phytopathologica 2
African journal of agricultural research 1
Agronomy 1
Agronomy Science and Biotechnology 1
Agronomy for Sustainable Development 1
Annual Review of Phytopathology 1
Australasian Plant Pathology 1
Basic and Applied Ecology 1
BioControl 1
Biochar 1
Chemical and Biological Technologies in Agriculture 1
Ciência Rural 1
Food Security 1
HortScience 1
Journal of Fungi 1
Journal of Plant Pathology 1
Nature Earth & Environment 1
PLOS ONE 1
Physiological and Molecular Plant Pathology 1
Plant Soil 1
Plants 1
Research, Society and Development 1
Sustainability 1
The Plant Pathology Journal 1
frontiers in Molecular Biosciences 1nrow(tab2)[1] 32set.seed(3)
old_par <- par(mar = c(0, 2, 0, 0), bg = NA)
p1 + wrap_elements(panel = ~wordcloud(words = tab2$journal, freq = tab2$n, min.freq = 1, max.words=200, random.order=FALSE, rot.per=0.25, colors=brewer.pal(6, "Dark2"))) + plot_annotation(tag_levels = "A")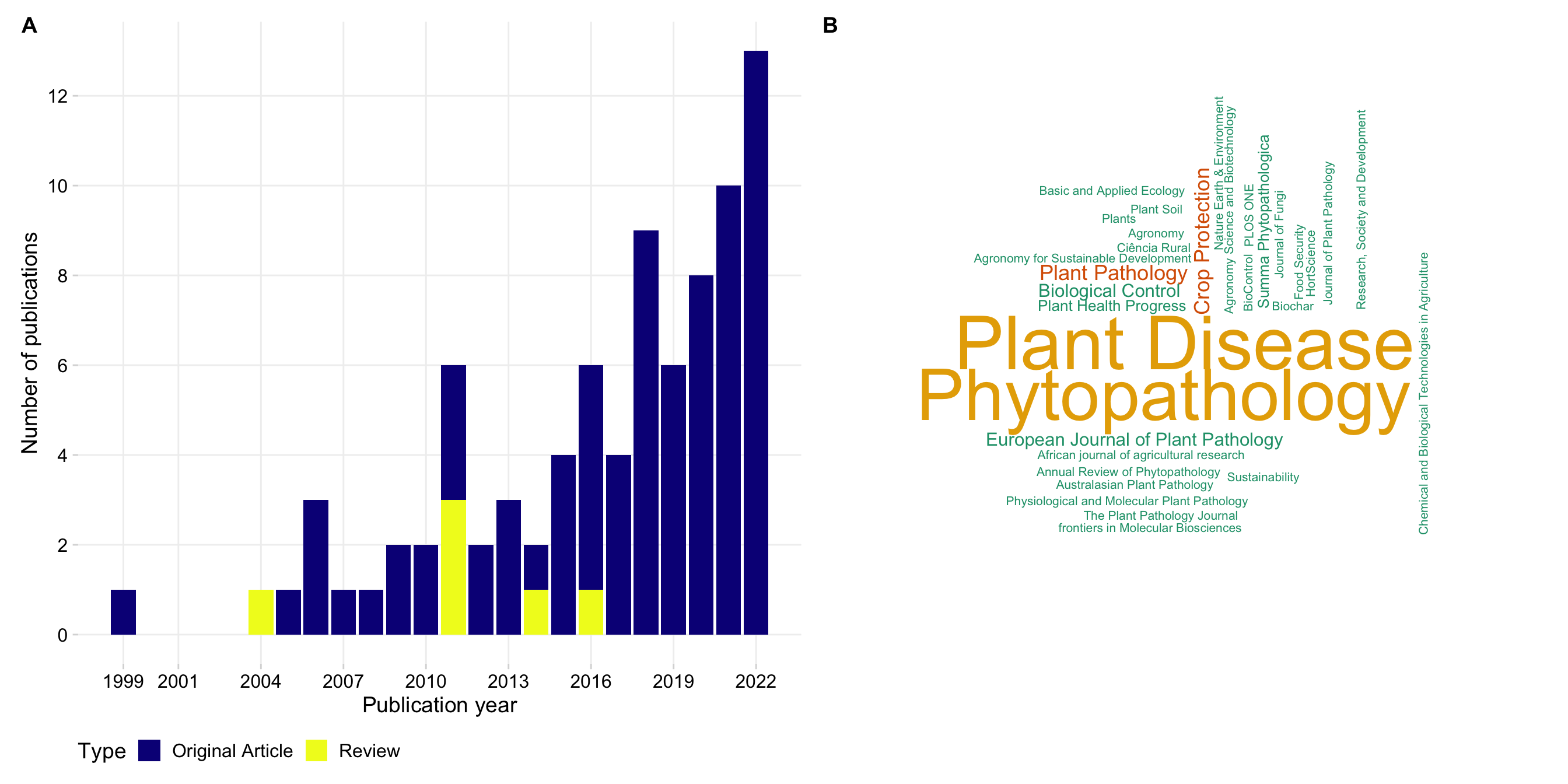
par(old_par)
ggsave("figs/figure1.png", width = 15, height = 8, bg = "white")Data characteristics
Source
p2 <- dat %>%
filter(article_type == "Original Article") %>%
tabyl(data_source) %>%
ggplot(aes(reorder(data_source, -n), n, fill = n))+
geom_col(fill = "#0d0887", width = 0.56)+
#geom_text(
# aes(x = data_source, y = n, label = n),
#position = position_dodge(width = 1),
#vjust = -0.5, size = 4) +
theme(legend.position = "bottom",
panel.grid.major=element_line(colour="grey94"))+
scale_y_continuous(breaks = c(0, 3, 6, 9, 12, 15, 18, 21, 24, 27, 30))+
labs(x = "Source of the data used in the analysis", y = "Number of original articles")
p2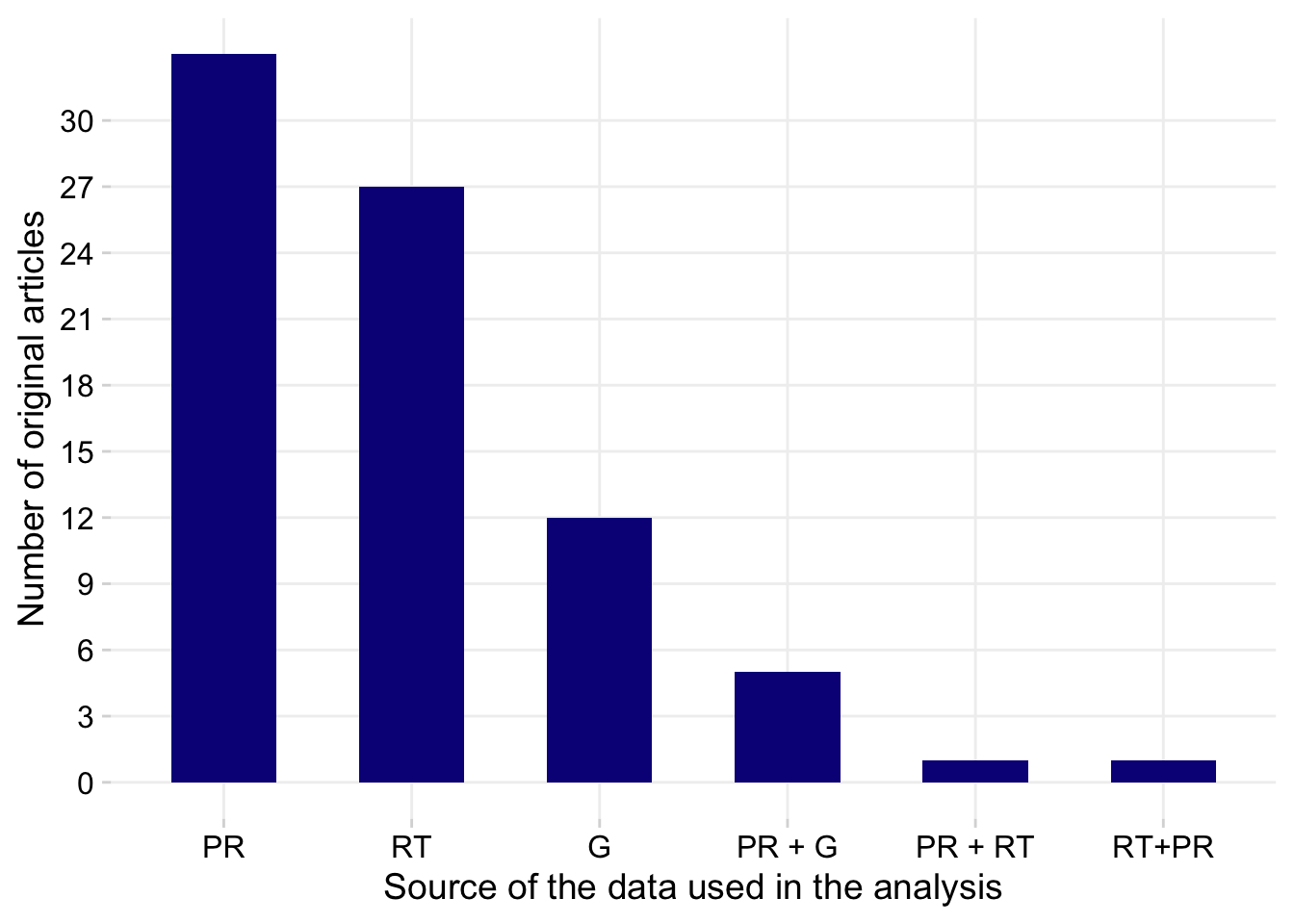
ggsave("figs/figure2.png", width =8, height = 6, bg = "white")Systematic review in PR?
dat |>
tabyl(systematic_review, data_source) systematic_review G PR PR + G PR + RT RT RT+PR NA_
no 11 0 0 0 27 1 0
yes 1 33 5 1 0 0 0
<NA> 0 0 0 0 0 0 6PRISM diagram?
dat |>
tabyl(sr_flow_diag) sr_flow_diag n percent valid_percent
no 71 0.83529412 0.8987342
yes 8 0.09411765 0.1012658
<NA> 6 0.07058824 NAStudy characteristics
Number of trials
dat |>
count(n_trials_total) |>
ggplot(aes(n_trials_total))+
geom_histogram(color = "white", fill = "#0d0887")+
labs(x = "N. of studies in the meta-analysis", y = "Frequency")+
scale_x_continuous(n.breaks = 10)+
theme(legend.position = "bottom",
panel.grid.major=element_line(colour="grey94"))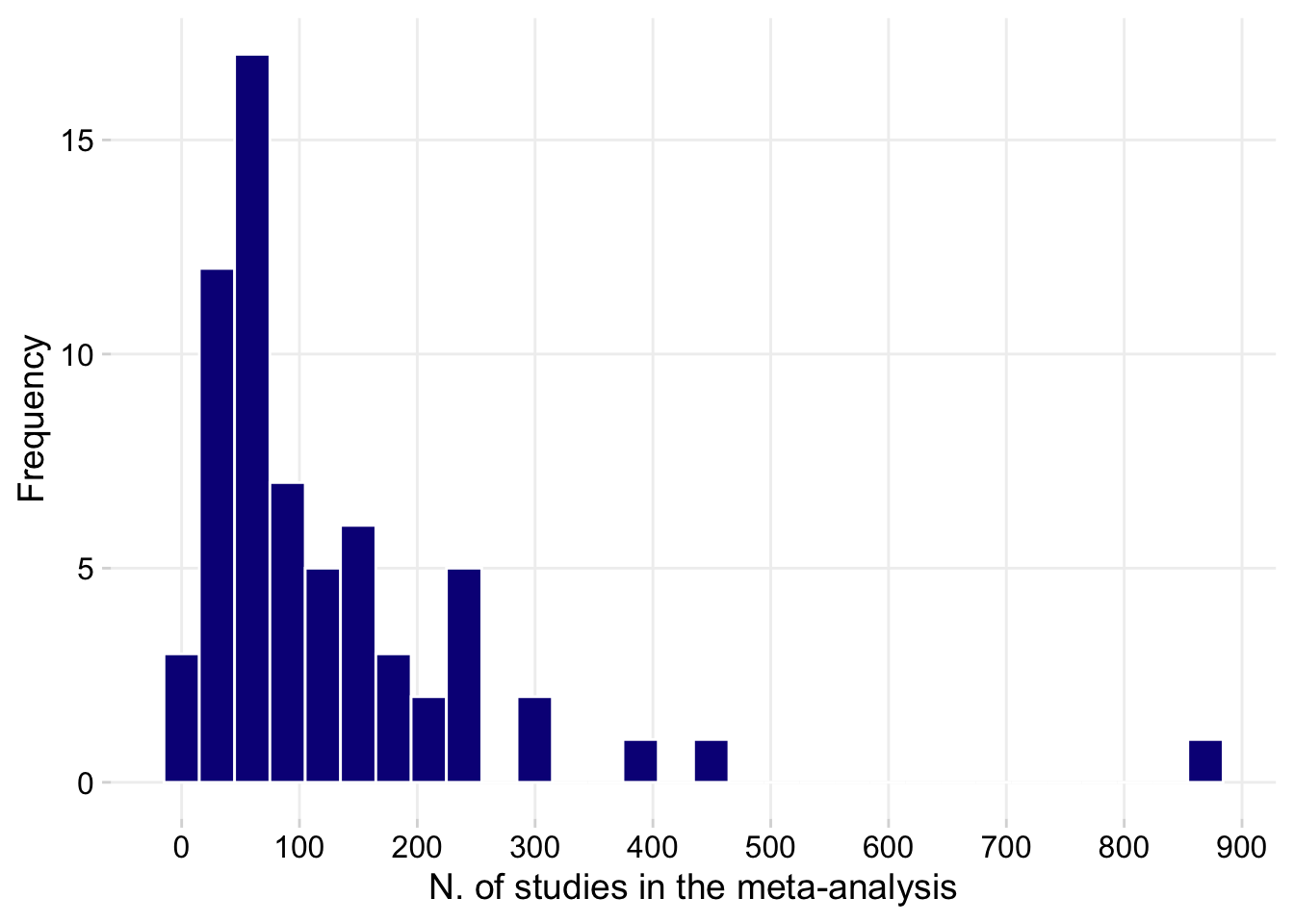
ggsave("figs/trials_study.png", width =6, height =4, bg = "white")
dat |>
count(n_trials_total) |>
summary() n_trials_total n
Min. : 10 Min. :1.000
1st Qu.: 46 1st Qu.:1.000
Median : 77 Median :1.000
Mean :121 Mean :1.288
3rd Qu.:162 3rd Qu.:1.000
Max. :879 Max. :6.000
NA's :1 By objective and product type
objective <- dat %>%
filter(article_type == "Original Article") %>%
tabyl(objective) |>
select(-percent)
type <- dat %>%
filter(article_type == "Original Article") %>%
filter(objective == "Product effects") %>%
tabyl(product_type) |>
select(-percent)
cbind(objective, type) objective n product_type n
1 Dis-toxin relationship 2 BCAs 12
2 Epidemic parameter 1 Bactericides 1
3 Host effects 1 Disinfestant 3
4 Management effects 8 Fertilizer 1
5 Monocyclic component 1 Fungicide 38
6 Product and host effects 1 Fungicide+BCAs 1
7 Product effects 58 Nematicide 1
8 Yld-Dis relationship 7 Sprayers and adjuvants 1Response variables
tab <- dat %>%
dplyr::select(response1 , response2, response3) %>%
pivot_longer(names_to = "type", values_to = "Variable", 1:3) %>%
select(Variable) %>%
filter(Variable != "NA") %>%
tabyl(Variable) %>%
select(-percent)
nrow(tab)[1] 38tab Variable n
100-seed weights 1
DON 6
Eggs/eggmass numbers in roots 1
RKN numbers in soil 1
Root galling 1
absolute yield 1
abundance 1
b^ 1
carpogenic germination 1
disease 1
galls index 1
incidence 15
incidence ratio 2
index 4
intensity 8
intercept 4
latent period duration 1
marketable yield 1
natural enemies 1
nematode density 1
number of cankers 1
number of eggs 1
number of galls 1
odds ratio 1
percent galled root 1
pest abundance 1
pests 1
plant biomass 2
plant growth 1
relative yield 1
sclerotia mass 1
seed germination 1
seedling emergence 1
severity 29
slope 4
test weight 1
toxin 1
yield 38library(wordcloud)
wordcloud(words = tab$Variable, freq = tab$n, min.freq = 1, max.words=200, random.order=FALSE, rot.per=0.25, colors=brewer.pal(5, "Dark2"))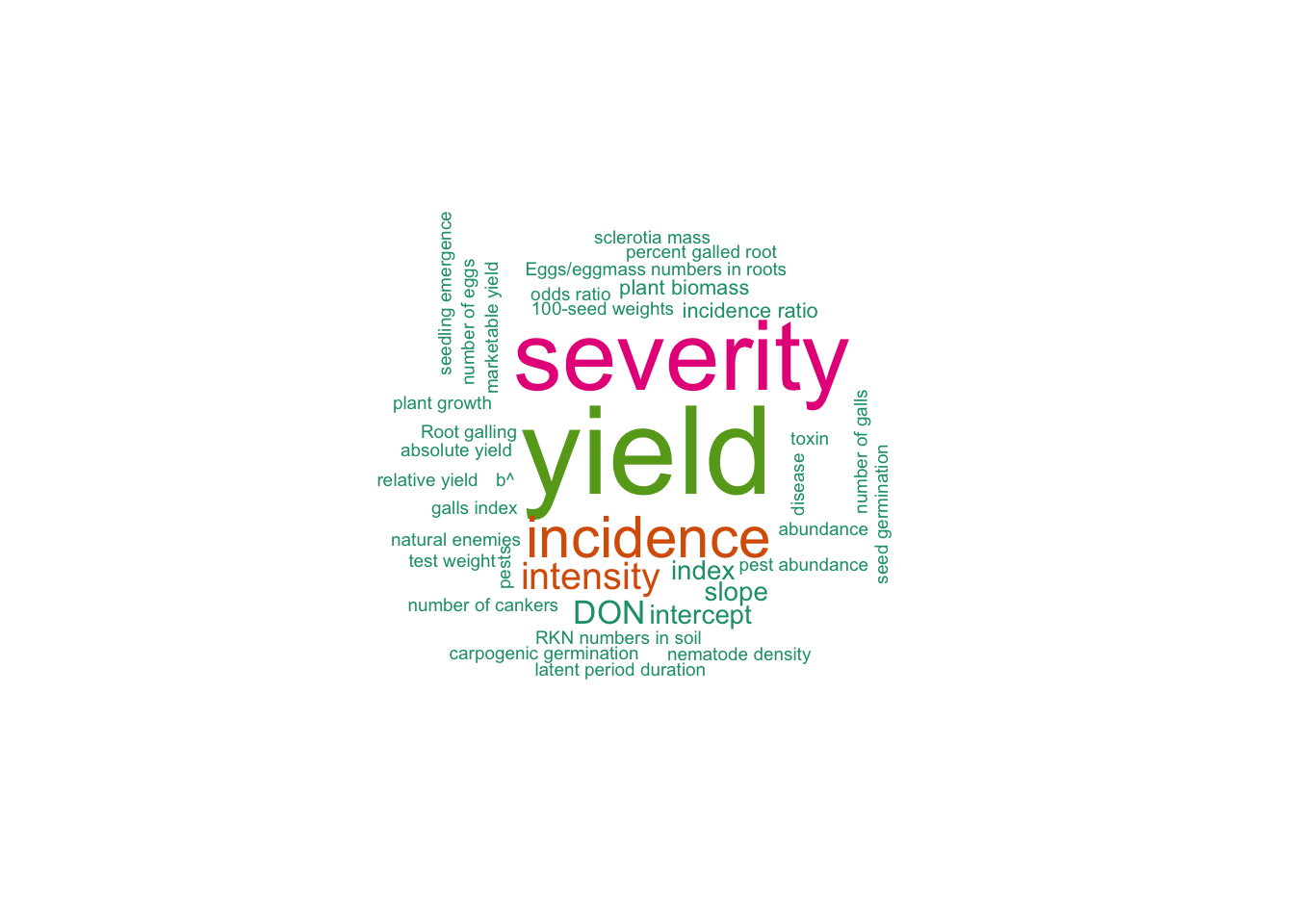
Number of responses per study
dat |>
tabyl(n_responses) n_responses n percent valid_percent
1 35 0.41176471 0.44871795
2 30 0.35294118 0.38461538
3 5 0.05882353 0.06410256
4 6 0.07058824 0.07692308
5 2 0.02352941 0.02564103
NA 7 0.08235294 NAMeta-analysis model characteristics
Effect sizes
es <- dat %>%
dplyr::select(effect_size_1, effect_size_2, effect_size_3, effect_size_4, effect_size_5) %>%
pivot_longer(names_to = "type", values_to = "value", 1:5) %>%
select(value) %>%
filter(value != "NA") %>%
tabyl(value) |>
adorn_totals()
es |>
arrange(-n) value n percent
Total 152 1.000000000
log ratio 36 0.236842105
means 35 0.230263158
log means 29 0.190789474
mean diff 12 0.078947368
slope 9 0.059210526
intercept 8 0.052631579
response ratio 8 0.052631579
Hedges' g 3 0.019736842
incidence ratio 3 0.019736842
odds ratio 3 0.019736842
Cohen's d 2 0.013157895
BPL b 1 0.006578947
log of d 1 0.006578947
r 1 0.006578947
risk ratio 1 0.006578947write_csv(es, "es.csv")Effect-size by common response variable
es <- dat %>%
dplyr::select(code, effect_size_1, effect_size_2, effect_size_3, effect_size_4, effect_size_5) %>%
pivot_longer(names_to = "type", values_to = "value", 2:6)
rv <- dat %>%
dplyr::select(code, response1 , response2, response3, response4, response5) %>%
pivot_longer(names_to = "type", values_to = "Variable", 2:6)
rv# A tibble: 425 × 3
code type Variable
<dbl> <chr> <chr>
1 1 response1 <NA>
2 1 response2 <NA>
3 1 response3 <NA>
4 1 response4 <NA>
5 1 response5 <NA>
6 2 response1 severity
7 2 response2 yield
8 2 response3 <NA>
9 2 response4 <NA>
10 2 response5 <NA>
# ℹ 415 more rowsrv2 <- left_join(es, rv, by = "code") |>
select(Variable, value) |>
filter(Variable %in% c("severity", "incidence", "yield", "intensity")) |>
tabyl(value, Variable)
rv2 value incidence intensity severity yield
Cohen's d 0 1 0 1
Hedges' g 0 3 0 0
incidence ratio 3 0 0 0
intercept 0 0 4 3
log means 4 0 17 17
log of d 0 0 1 0
log ratio 14 0 18 19
mean diff 1 1 2 9
means 11 2 11 27
odds ratio 3 0 0 0
r 0 0 1 0
response ratio 0 1 2 7
risk ratio 0 1 0 0
slope 0 0 5 4
<NA> 39 31 84 118write_csv(rv2, "es2.csv")Sampling variance
dat |>
tabyl(`Inverse variance`) Inverse variance n percent valid_percent
no 4 0.04705882 0.05063291
not reported 4 0.04705882 0.05063291
yes 71 0.83529412 0.89873418
<NA> 6 0.07058824 NAHeterogeneity test
dat |>
tabyl(`Heterogenity test`) Heterogenity test n percent valid_percent
H2 and I2 1 0.01176471 0.01923077
I2 9 0.10588235 0.17307692
I2 and R2 1 0.01176471 0.01923077
LRT 8 0.09411765 0.15384615
LRT and I2 1 0.01176471 0.01923077
LRT and R2 2 0.02352941 0.03846154
Q 8 0.09411765 0.15384615
Q and I2 4 0.04705882 0.07692308
Q, H2 and I2 1 0.01176471 0.01923077
Q, I2 5 0.05882353 0.09615385
R2 2 0.02352941 0.03846154
Wald 8 0.09411765 0.15384615
Z? 1 0.01176471 0.01923077
no 1 0.01176471 0.01923077
<NA> 33 0.38823529 NAEstimator
dat |>
tabyl(estimator) estimator n percent valid_percent
DL 12 0.14117647 0.1518987
ML 56 0.65882353 0.7088608
Unknown 11 0.12941176 0.1392405
<NA> 6 0.07058824 NAGeneral approach
dat |>
tabyl(ma_approach) ma_approach n percent valid_percent
frequentist 76 0.89411765 0.96202532
frequentist and Bayesian 3 0.03529412 0.03797468
<NA> 6 0.07058824 NAMA basic model
dat |>
tabyl(ma_model) ma_model n percent valid_percent
MTC 23 0.27058824 0.29113924
Single treatment 53 0.62352941 0.67088608
not applicable 3 0.03529412 0.03797468
<NA> 6 0.07058824 NAdat |>
filter(ma_model == "MTC") |>
tabyl(effect_size_1, effect_size_2) effect_size_1 intercept log means log ratio mean diff means NA_
log means 0 4 0 0 8 1
log ratio 0 0 4 0 0 0
mean diff 0 0 0 1 0 1
means 1 0 0 0 1 2MA model n. of effects
dat |>
tabyl(ma_model_2) ma_model_2 n percent valid_percent
Kruskall-Wallis 1 0.01176471 0.01265823
fixed- and random-effects 4 0.04705882 0.05063291
fixed-effects 7 0.08235294 0.08860759
linear regression 1 0.01176471 0.01265823
mixed-effects 33 0.38823529 0.41772152
non-parametric 1 0.01176471 0.01265823
not informed 1 0.01176471 0.01265823
random- and mixed-effects 3 0.03529412 0.03797468
random-effects 27 0.31764706 0.34177215
random-effects and mixed-effects 1 0.01176471 0.01265823
<NA> 6 0.07058824 NANumber of variables
dat |>
tabyl(ma_n_variables) ma_n_variables n percent valid_percent
bivariate 1 0.01176471 0.01265823
univariate 78 0.91764706 0.98734177
<NA> 6 0.07058824 NAModerator analysis?
dat |>
tabyl(moderator) moderator n percent valid_percent
no 12 0.14117647 0.1518987
yes 67 0.78823529 0.8481013
<NA> 6 0.07058824 NAdat |>
tabyl(moderator_model) moderator_model n percent valid_percent
covariate 6 0.07058824 0.08955224
covariate and metaregression 3 0.03529412 0.04477612
metaregression 2 0.02352941 0.02985075
subgroup 31 0.36470588 0.46268657
subgroup and metaregression 23 0.27058824 0.34328358
subroup 2 0.02352941 0.02985075
<NA> 18 0.21176471 NASoftware characteristics
General software
software <- dat |>
tabyl(general_software) |>
arrange(general_software)
software general_software n percent valid_percent
ARM ST 1 0.01176471 0.0125
CMA 5 0.05882353 0.0625
GENSTAT 1 0.01176471 0.0125
MetaWin 3 0.03529412 0.0375
OpenMee 1 0.01176471 0.0125
R 29 0.34117647 0.3625
SAS 31 0.36470588 0.3875
Stata 1 0.01176471 0.0125
WinBUGS 1 0.01176471 0.0125
not mentioned 2 0.02352941 0.0250
not reported 5 0.05882353 0.0625
<NA> 5 0.05882353 NAwrite_csv(software, "software.csv")dat |>
tabyl(MA_software) MA_software n percent valid_percent
CMA 4 0.04705882 0.06153846
N.M. 1 0.01176471 0.01538462
PROC GLIMMIX 8 0.09411765 0.12307692
PROC MIXED 20 0.23529412 0.30769231
PROC UNIVARIATE 1 0.01176471 0.01538462
SAS macros 2 0.02352941 0.03076923
brms 1 0.01176471 0.01538462
lme4 2 0.02352941 0.03076923
lme4 anc R2jags 1 0.01176471 0.01538462
metafor 25 0.29411765 0.38461538
<NA> 20 0.23529412 NAData summary
Results in table?
dat |>
tabyl(res_table) res_table n percent valid_percent
no 9 0.10588235 0.1139241
yes 70 0.82352941 0.8860759
<NA> 6 0.07058824 NAResults in plot for raw data
dat |>
tabyl(res_plot_raw) res_plot_raw n percent valid_percent
Yes 1 0.01176471 0.01265823
no 43 0.50588235 0.54430380
yes 35 0.41176471 0.44303797
<NA> 6 0.07058824 NAResult in forest plot
dat |>
tabyl(res_forest) res_forest n percent valid_percent
no 64 0.75294118 0.8101266
yes 15 0.17647059 0.1898734
<NA> 6 0.07058824 NAEconomic analysis
dat |>
tabyl(econ_analysis) econ_analysis n percent valid_percent
no 60 0.70588235 0.7594937
yes 19 0.22352941 0.2405063
<NA> 6 0.07058824 NA